NIEHS Scientists Redirect Efforts to Fight Covid-19
4th article in a series on intramural NIH scientists who pivoted their research project to pursue a pandemic-related issue.
At the beginning of the Covid-19 pandemic, NIEHS scientists Dr. Mario Borgnia and Dr. Robin Stanley began contributing to NIH’s research efforts against SARS-CoV-2. They used their experience in structural biology to learn more about the proteins involved in the essential function of the virus that causes Covid-19.
Giving Serendipity a Boost
Borgnia said he had a choice: he could sit on his hands or do something to increase the probability of serendipitously finding something that would help researchers learn about SARS-CoV-2.
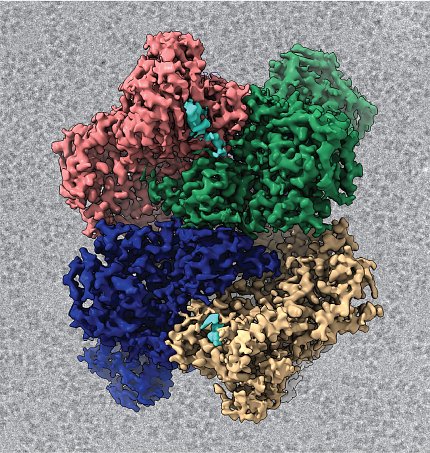
Photo: MEREDITH FRAZIER
“I wasn’t going to sit at home and be depressed,” said Borgnia, director of the Molecular Microscopy Consortium at NIEHS. “I participated in the Covid-19 scientific interest group and helped organize talks so people had something constructive to do.”
The consortium allows scientists to use single-particle cryogenic-electron microscopy (cryo-EM) and other tools employing high-tech microscopes to decipher macromolecular structures at the atomic level.
Cryo-EM is an advanced microscopy methodology to image complex biological molecules like proteins.
To obtain the images, molecules are frozen in a thin layer of ice on a grid. Then, they are bombarded with a beam of electrons to produce projection images. Next, images are combined to construct a 3-D map of molecules. The resulting images are like CAT scans for molecules.
The week before most NIH employees and trainees began teleworking full-time, Borgnia’s team began to study the protein spikes conferring the corona-like appearance to the SARS-CoV-2 virus. These spikes bind to the surface of cells, the first step before the virus can invade a host cell.

Photo: Steve McCaw
“We concentrated our efforts on one particular set of proteins rather than a wider range of proteins,” he said. “We were well positioned to get started and just put all of our firepower on the spike of the coronavirus.”
Once he had the spike protein’s structure, he began collaborating with researchers who were developing antibodies that would neutralize the spike. Essentially, his consortium created a pipeline so researchers could study how the spike behaves when exposed to different molecules.
Studying the SARS-Cov-2 virus wasn’t a hard transition for him. Earlier in his career, Borgnia studied a protein, Env, the envelope glycoprotein, found on the surface of the HIV virus. Env allows the virus to target and attach to specific cell types.
Borgnia has done most of his work remotely from home. His team gathers via Zoom twice each day for check-in and check-out meetings that give structure to a remote work day. They have accelerated the transition to remote operation of the microscope, which allowed them to increase productivity both on-site and away from the workplace. During this period, a member of his team would go into the lab to prepare a sample to be imaged and then leave. Team members could then control the microscope from the sheltered environments of their homes.
While the pandemic has been terrible, it has given Borgnia the opportunity to collaborate with researchers around the world who are studying the virus.
“My ability to communicate has increased dramatically,” he said. “I can now sit in the same virtual room with people who are all over the world. I don’t have to travel and can spend time talking to people.”
‘I Need to Be in a Lab’

Photo: MEREDITH FRAZIER
Stanley never planned on studying the virus that causes Covid-19 because she didn’t know much about viruses. Since she started her lab in 2014, Stanley has studied RNA processing enzymes involved in ribosome assembly and tRNA maturation. Shortly after she began teleworking full time, she realized she didn’t like it. She decided to pivot from studying pre-rRNA processing to viral RNA processing because scientists working on Covid-19 research are allowed in labs.
“I don’t like working at home. I need to be in the lab,” said Stanley, Earl Stadtman investigator and head of the NIEHS nucleolar integrity group. “I gave myself a crash course in the coronavirus and studied the different viral proteins. There was 1 viral protein that actually shared an awful lot of similarities with 2 RNA processing enzymes that we work on in our lab.”
In addition, she thought the experience would be a great learning opportunity for her trainees. “It gave the lab something to focus on during that crazy period of time when we didn’t know what was going on,” she said.

Photo: Steve McCaw
Stanley applied to the Intramural Targeted Anti-Covid-19 funding program, which provided $12 million to intramural investigators for research into understanding and/or combatting the virus. She was awarded funding to study nonstructural protein 15 (Nsp15), which is an RNA processing enzyme found in all coronaviruses. Without Nsp15, the virus becomes vulnerable to the host’s immune system. It’s thought that the protein helps the virus evade detection.
Although Nsps do not become part of the mature virus, they “are really, really important for the virus,” she explained. They are involved in virus replication, virus assembly and the evasion of host viral sensors.
Nsp15 is an endoribonuclease. It’s an enzyme that cuts RNA in two—like a pair of molecular scissors. Some nucleases cut anything; others look only for a very specific sequence. Nsp15 only cuts after uridines, one specific nucleotide of RNA.
She collaborated with Borgnia to use cryo-EM to visualize the protein at a molecular level. These images have provided lots of information about why Nsp15 behaves the way it does.
“The big question we wanted to answer: how does it recognize and cut viral RNA targets?” she said. “Thankfully, through a combination of biochemistry, cryo-EM and some other techniques, we’ve been sort of able to answer that question. There are always more questions that arise from research, but we’ve been able to visualize the protein bound to some RNA, which has provided an abundance of information.”
At the beginning of her research, she thought making the protein would be easy. It turned out to be an unexpected challenge. The method Stanley typically uses to express proteins didn’t work. Normally, her lab hijacks a bacteria’s ability to make protein.
“But it turns out that bacteria don’t like that protein, because likely it’s toxic. When you express the nuclease in bacteria, it’s probably chewing up all the bacteria’s RNA and it’s very unhappy,” she explained.
After a month of trial and error, her group overcame this limitation and figured out another way to make the protein. Hopefully, these findings can be used to provide a structural framework for the development of new therapeutics.
Right now, her lab has begun to focus attention back on projects predating the pandemic. Two members are still working on NSP15.
“The ability to pivot is one of the awesome things about working at NIH,” she concluded. “Hopefully, a pandemic won’t happen again, but, if it does happen in the future, researchers at NIH can do it again.”
