Drivers and Passengers
Chanock Explores Genetics of Cancer Susceptibility

Great teachers inspire as they impart wisdom. Such was the case during a recent lecture by NCI’s Dr. Stephen Chanock, who managed to fit his professional journey, career advice, a history lesson and decades of research into a captivating hour.
“Trust your instincts and follow your interests, wherever they take you,” advised Chanock, who called his professional path a tale of two long woods. He began his NCI career 30 years ago, though his connection to NIH started much earlier.
Chanock grew up on Longwood Drive near the Bethesda campus and fondly recalls visiting NIH with his father, who worked at NIAID. “Bldg. 7…was a standard Saturday morning playground for me as a young child,” he said.
Initially, Chanock didn’t plan to study medicine. His interests first took him to study music in college. But then he wound up on Longwood Ave., in Boston, Mass., studying medicine at Harvard.

His professional career would also take a turn. Chanock arrived at NIH in 1991 to study oncology and infectious diseases but his interests pivoted by 2000 as genomics research gained momentum. Mentors had told him to pursue the interesting questions, which for him turned out to be, not the section where he was a tenured investigator, but the Division of Cancer Epidemiology and Genetics, where he is currently the director.
Speaking at a virtual Clinical Center Grand Rounds Great Teachers talk, Chanock described some of the research his genetics lab has conducted into the biology, heritability and risks of cancer.

Photo: Ernie Branson
The Human Genome Project opened a whole new world of discovery. “The excitement of characterizing and sequencing nearly all 3.1 billion bases,” said Chanock, “gives us the opportunity to over and over look both at the germline and the tumors to understand what kinds of changes we see—what changes are drivers and what changes may be passengers—due to the cancer phenotype being expressed.”
Medical observations of cancer running in families began more than a century ago, before anyone had ever described a gene. Then came a game-changer, in 1969, when research by two NCI scientists—Drs. Fred Li and Joseph Fraumeni—launched the era of cancer genetics.
“This really changed the landscape, the capacity to see that different types of cancer could arise in families with supposedly the same genetic makeup,” said Chanock.
He has devoted his career to the genetic underpinnings of cancer, from identifying mutations to understanding genetic-environmental interactions that make some people more susceptible to cancer. Most attributable risks, he said, come from environmental factors such as smoking, obesity and exposure to toxic chemicals and radiation.
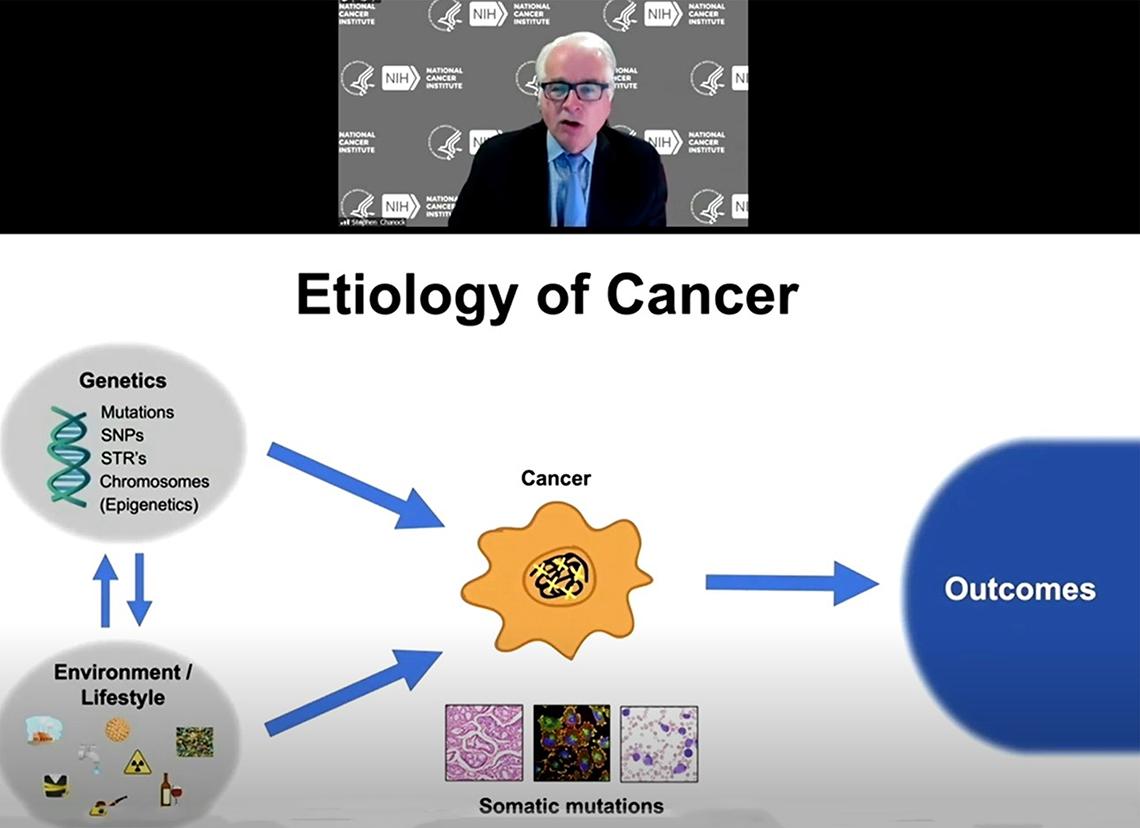
“These activities, these environmental/lifestyle choices—or sometimes they’re not choices, but exposures—interact with the host genome,” he said. “That relationship is really the central question I see that the next 20 years of cancer research has to investigate thoroughly to understand the fundamentals that would better enable us to come up with both precision preventive models and also therapies.”
Environmental stimuli lead to mutations in the cells. Cancer develops progressively over time, as different driver mutations take over, explained Chanock.
“Somewhere between the exposures of lifestyle and baseline genetics, our genetic germline is indeed falling apart,” he said. His lab continues to isolate variations in large population and family studies, trying to understand not only the interaction between exposures and mutations, but also what happens in between.

Photo: NCI/istock
One large, longterm study exploring the interaction between genetics and environmental exposures is looking at the effects of radiation exposure from the 1986 Chernobyl nuclear accident in Ukraine. One recent finding, said Chanock, is that in people younger than 18 at the time of exposure who developed cancers over the next 20 years, nearly 95 percent of the mutations involved a specific pathway, MAPK. A separate family study is looking at whether direct radiation exposure in the cleanup workers could affect successive generations. Thankfully, the data do not support a strong transgenerational effect. This has important implications for those exposed after the Chernobyl accident but also those exposed after the 2011 Fukushima nuclear accident in Japan.
Looking at large populations through genome-wide association studies (GWAS) helps investigators stratify cancer risk. To date, investigators can validate and fully understand only 5-10 percent of all GWAS signals. They do know there are more than 1,200 separate regions across the genome associated with 1 or more, and up to 30, cancers, said Chanock. But it takes years to understand and ascribe functions to each one of these specific GWAS loci.
“What’s also striking is that only a handful of these 1,200 [regions] have a strong signal that’s associated with outcome,” he said. “So what we’re looking at is this complex temporal model of cancer. What kinds of perturbations set the stage for cancer to develop in an individual may be very different than what’s responsible for either protecting or not protecting an individual as they undergo therapy and try to control cancer that’s growing out of control.”
Further complicating the landscape, many variants each contribute a small effect to a large fraction of cancers. And each cancer has a different genetic architecture. About 85 percent of variants are specific to a certain cancer. With only 15 percent of variants shared between cancers, different strategies are needed to target different cancers.
Advances in cancer research rely on pooling dozens of large, distinct GWAS studies from around the world. “[These studies allow us] to work forwards and backwards to keep getting bits and pieces or handles on what we see as the complex paradigm of development of cancer,” Chanock said. “The value of data-sharing couldn’t be overstated, because it’s really team science at its best.”
GWAS helps investigators assign polygenic risk scores with the goal of using the data for clinical care. Looking within a population, Chanock and colleagues examine all the SNPs (“snips”—or, single nucleotide polymorphisms—which are variations at a single point in DNA) associated with risk for a particular cancer that has high heritability. “That helps us to have a risk score with a predictive ability,” he said.
In a study of more than 100,000 people, for example, investigators found higher genetic risk profiles for prostate cancer in men of African-American ancestry. By assigning risk scores to populations of different descents, researchers can better understand unique biologic features that could lead to better therapies and better opportunities to stratify groups of individuals at high risk for more intensive screening.
For breast cancer, the deleterious mutations in the BRCA 1 and 2 genes explain less than a fifth of cases in the U.S., said Chanock. GWAS data has helped piece together another 40 percent of breast cancer-causing genes, but a third of cases remain unexplained. Large, multi-ancestral GWAS are helping researchers understand subtypes and discover variants to improve treatment, prevention and survival.
“Don’t be afraid to look at old problems with new approaches,” advised Chanock, who reiterated the importance of team science. “Context matters. Numbers matter even more. Your colleagues matter most.”
